Nanoscopic DNA pyramids that change shape when sent different chemical signals, have been demonstrated by researchers in the UK and Germany. Such structures could act as the motors of nanoscale robots, they say.
The researchers demonstrate the operation of reconfigurable DNA tetrahedra whose shapes change precisely and reversibly in response to specific molecular signals. Shape changes are confirmed by gel electrophoresis and by bulk and single-molecule Förster resonance energy transfer measurements. DNA tetrahedra are natural building blocks for three-dimensional construction9; they may be synthesized rapidly with high yield of a single stereoisomer, and their triangulated architecture conveys structural stability. The introduction of shape-changing structural modules opens new avenues for the manipulation of matter on the nanometer scale.
The DNA piston work is described at New Scientist magazine.
Other researchers have previously built DNA devices capable of walking along proteins or functioning like nanoscopic robot arms, but precise control of these 3D structures has proven difficult.
Now Andrew Turberfield of Oxford University in the UK, and colleagues at the University of Bielefeld in Germany, have shown how carefully crafted DNA structures can be made to self assemble and change shape when sent specific DNA signals.
The researchers built tetrahedrons – structures with four triangular sides – using four short DNA “struts” that join at each end. The process exploits the way DNA is held together by complementary bases that form the rungs of a ladder-like structure.
Tuberfield and colleagues first created DNA molecules with some bases on one side of the “ladder” left exposed. These bases were carefully chosen to match those of other DNA molecules so that, when mixed together, the right combination of DNA strands assembled into a tetrahedron.
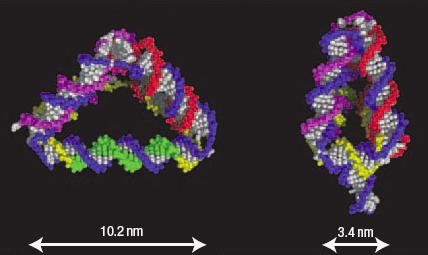
The same trick can change the tetrahedron shape, by causing the struts to extend or shorten. This is possible if a strut has a loop of excess DNA in its middle (see image, top right). Adding a “fuel” strand of DNA straightens out the loop by binding onto it, and makes the strut extend.
A different sequence of “anti-fuel” DNA returns the strut to its normal length. “The fuel is designed to have a dangling free end,” explains Turberfield. “The anti-fuel eventually displaces the fuel, allowing the loop to reform.” The process can change the length of a strut from 3.4 nanometres in length 10.2 nm and back again.
In experiments, the researchers made cages with two extendible struts that could be independently controlled using different DNA sequences. In theory, it should be possible to create cages in which every strut can be controlled independently, Tuberfield says.
The researchers are now exploring ways to build larger structures using tetrahedral DNA structures as building blocks.
Drug delivery
In a commentary article concerning the new research, Chengde Mao at Purdue University
, West Lafayette, Indiana, US, who was not involved with the work, says the structures could perhaps be used to carry a payload and release it on demand.
“It is likely that these studies will lead to responsive molecular cages that can encapsulate guests and release them on demand,” he writes. “For example, it is now possible to enclose proteins in a DNA tetrahedron.” The cages can also be decorated with carefully placed proteins, perhaps providing a way to get the body to treat cages in a particular way.
Turberfield agrees: “Using the cages to hold and release proteins could have uses in drug delivery.” For example, he says, drug compounds normally destroyed by the body could travel to a particular organ in a DNA cage, before being released.
FURTHER READING
Andrew Turberfield of Oxford University site

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.