Scientists develop a catalog of 1000 tumors with their alterations and drug sensitivities
There are thousands of scientific papers dedicated to a particular type of tumor, a particular gene, a type of specific molecular lesion or the effect of a particular drug. However, there are very few examples of publications that integrate these four concepts (type of tumor, gene alteration and drug) in a significant amount of samples. An article published in Cell, in collaboration with the group of Dr. Manel Esteller, Director of the Epigenetics and Cancer Biology Program at the Bellvitge Biomedical Research Institute (IDIBELL), ICREA researcher and Professor of Genetics at the University of Barcelona, provides us with this important source of information.
The researchers started from 1,000 cancer cell lines derived from 29 different cell types and different organs to simultaneously obtain their genetic alterations (mutations and copy number of genes), epigenetic alterations (DNA methylation) and expression alterations, confronting them with their different sensitivity to 265 antitumor drugs. It should also be noted that the results were validated in 11,000 additional human tumor samples obtained from surgical removals.
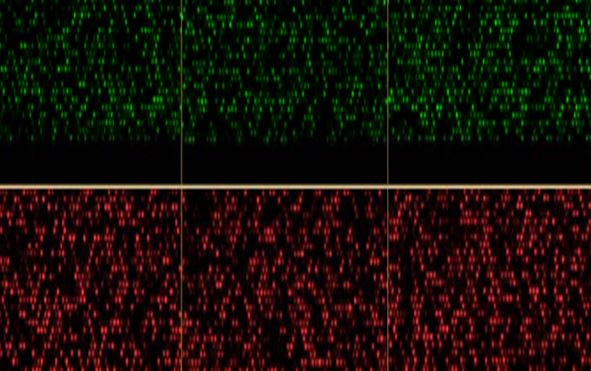
Example of an epigenome derived from a tumor sample included in study. Green fluorescence signals indicate a loss of epigenetic signal, while red indicates a gain of epigenetic signal. Nearly half a million points of epigenetic information in the human g
Journal Cell – A Landscape of Pharmacogenomic Interactions in Cancer
Highlights
• We integrate heterogeneous molecular data of 11,289 tumors and 1,001 cell lines
• We measure the response of 1,001 cancer cell lines to 265 anti-cancer drugs
• We uncover numerous oncogenic aberrations that sensitize to an anti-cancer drug
• Our study forms a resource to identify therapeutic options for cancer sub-populations
Summary
Systematic studies of cancer genomes have provided unprecedented insights into the molecular nature of cancer. Using this information to guide the development and application of therapies in the clinic is challenging. Here, we report how cancer-driven alterations identified in 11,289 tumors from 29 tissues (integrating somatic mutations, copy number alterations, DNA methylation, and gene expression) can be mapped onto 1,001 molecularly annotated human cancer cell lines and correlated with sensitivity to 265 drugs. We find that cell lines faithfully recapitulate oncogenic alterations identified in tumors, find that many of these associate with drug sensitivity/resistance, and highlight the importance of tissue lineage in mediating drug response. Logic-based modeling uncovers combinations of alterations that sensitize to drugs, while machine learning demonstrates the relative importance of different data types in predicting drug response. Our analysis and datasets are rich resources to link genotypes with cellular phenotypes and to identify therapeutic options for selected cancer sub-populations.
“The study has been led by the Sanger Institute in the UK, which are global pioneers in the genetic study of tumors. They asked us to analyze the epigenome of thousands of described samples and so we did. It was a though job, but the result was worth it: we now have a comprehensive map of genetic and epigenetic lesions of human tumors that serve to predict responses to a wide range of cancer drugs “- says Dr. Manel Esteller, co-author of the Cell paper – “moreover, all the obtained information has been released under open access licensing to be freely available to all researchers in the world. This way, if someone has a particular interest in a gene for a type of tumor or a drug he could quickly look it up, even view the relationships between them. “
SOURCES- Journal Cell, Bellvitge Biomedical Research Institute (IDIBELL)

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.