From nanowerk, Harvard and Princeton researchers have developed steps toward biological computers These machines can convert cellular signals and activity into easily detectable markers.
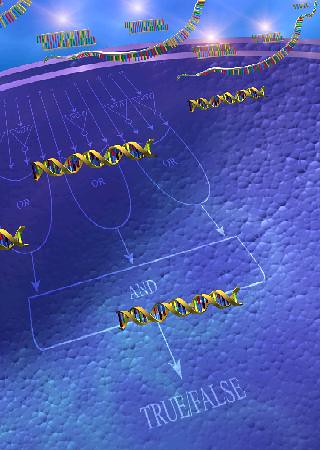
1. The entire computer (device able to perform boolean logic) is “encoded” by genes made of engineered DNA molecules.
2.The genes enter the cell using special delivery techniques and interact with the existing cellular activity.
3. Components of the computer, generated from its genes, are short hairpin RNA molecules known as shRNA and messenger ribonucleic acid molecules known as mRNA.
4. Both mRNA molecules generate the same output protein. shRNA molecules are designed to target and destroy the mRNA molecules using short color-coded mRNA stretches (sequences). They “hitchhike” on the biological mechanism known as RNA interference (RNAi).
5. Dicer enzyme, present in the cell, binds to shRNA and cleaves the loop-shaped tail
6. Another cellular enzyme, called RISC, binds to the dicer-processed shRNA and unwinds it, capturing one of the two strands
7. RISC bound to the RNA strand may now bind to the target region in the mRNA. The nucleotide sequence of the RNA strand guides RISC to its destination
8. Combination of RISC activity and other cellular mechanisms lead to the degradation of the mRNA.
9. All three shRNAs are engaged in a similar process, and each of them targets a different region within mRNA molecules, as indicated by black blunt-end arrows
10. Ultimately, any logic algorithm may be implemented using appropriate extensions.
Evaluating Boolean logic equations inside cells, these molecular automata will detect anything from the presence of a mutated gene to the activity of genes within the cell. The biocomputers’ “input” is RNA, proteins and chemicals found in the cytoplasm; “output” molecules indicating the presence of the telltale signals are easily discernable with basic laboratory equipment.
“Currently, we have no tools for reading cellular signals,” Yaakov Benenson says. “These biocomputers can translate complex cellular signatures, such as activities of multiple genes, into a readily observed output. They can even be programmed to automatically translate that output into a concrete action, meaning they could either be used to label a cell for a clinician to treat or they could trigger therapeutic action themselves.”
Benenson and his colleagues demonstrate in their Nature Biotechnology paper that biocomputers can work in human kidney cells in a culture. Research into the system’s ability to monitor and interact with intracellular cues such as mutations and abnormal gene levels is still in progress.
Further info:
Here is animation of the molecular automata

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.

