Harvard has a new protein array lab on a chip. The system could produce any desired protein from synthesized DNA placed on the chip. It is another step in making protein production and engineering and biotechnology in general faster and cheaper.
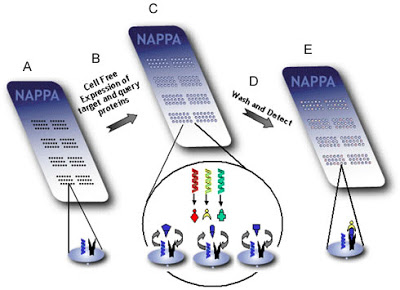
Schematic representation of screening protein-protein interactions with NAPPA.
(A) On each spot, a target plasmid and an affinity capture molecule are linked to the prepared glass slide.
(B) The slide is bathed with the cell-free in vitro transcription/translation mix, containing one or more query plasmids expressing different affinity tags.
(C) The target proteins are expressed and immobilized on the spots, as are query proteins if they bind to that target.
(D) The wells are then washed to remove unbound protein.
(E) Target-query complexes are detected by fluorescence.
Existing protein arrays involve the tedious and lengthy process of expressing proteins in living cells followed by purifying, stabilizing, and spotting the samples. This process is a bottleneck in the preparation of the arrays. Moreover, functionally active proteins require careful manipulation, and the less that is needed the better. Our approach to developing a protein array, a Nucleic Acid-Programmable Protein Array (NAPPA), replaces the complex process of spotting purified proteins with the straightforward and much simpler process of spotting plasmid DNA.

NAPPA Microarray. Genes (35) involved in eukaryotic DNA replication are expressed and immobilized in situ as GST fusion proteins on a microarray format. Expression of target proteins was detected by fluorescence. The samples were arrayed using a GMS 417 pin arrayer with 900µm spacing.

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.