Researchers from Northwestern University and Yale University have developed a user-friendly technology to help scientists understand how proteins work and how to fix them when they are broken. Such knowledge could pave the way for new drugs for a myriad of diseases, including cancer.
The human body has a nifty way of turning its proteins on and off to alter their function and activity in cells: phosphorylation, the reversible attachment of phosphate groups to proteins. These “decorations” on proteins provide an enormous variety of function and are essential to all forms of life. Little is known, however, about how this dynamic process works in humans.
Using a special strain of E. coli bacteria, the researchers have built a cell-free protein synthesis platform technology that can manufacture large quantities of these human phosphoproteins for scientific study. This will enable scientists to learn more about the function and structure of phosphoproteins and identify which ones are involved in disease.
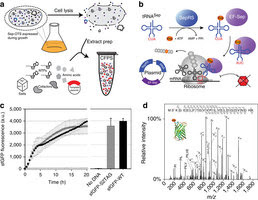
CFPS platform with an expanded genetic code for the production of phosphoproteins.
Trouble in the phosphorylation process can be a hallmark of disease, such as cancer, inflammation and Alzheimer’s disease. The human proteome (the entire set of expressed proteins) is estimated to be phosphorylated at more than 100,000 unique sites, making study of phosphorylated proteins and their role in disease a daunting task.
“Our technology begins to make this a tractable problem,” Jewett said. “We now can make these special proteins at unprecedented yields, with a freedom of design that is not possible in living organisms. The consequence of this innovative strategy is enormous.”
Jewett, associate professor of chemical and biological engineering at Northwestern’s McCormick School of Engineering, and his team worked with Yale colleagues led by Jesse Rinehart. Jewett and Rinehart are co-corresponding authors of the study.
As a synthetic biologist, Jewett uses cell-free systems to create new therapies, chemicals and novel materials to impact public health and the environment.
“This work addresses the broader question of how can we repurpose the protein synthesis machinery of the cell for synthetic biology,” Jewett said. “Here we are finding new ways to leverage this machinery to understand fundamental biological questions, specifically protein phosphorylation.”
Jewett and his colleagues combined state-of-the-art genome engineering tools and engineered biological “parts” into a “plug-and-play” protein expression platform that is cell-free. Cell-free systems activate complex biological systems without using living intact cells. Crude cell lysates, or extracts, are employed instead.
Specifically, the researchers prepared cell lysates of genomically recoded bacteria that incorporate amino acids not found in nature. This allowed them to harness the cell’s engineered machinery and turn it into a factory, capable of on-demand biomanufacturing new classes of proteins.
“This manufacturing technology will enable scientists to decrypt the phosphorylation ‘code’ that exists in the human proteome,” said Javin P. Oza, the lead author of the study and a postdoctoral fellow in Jewett’s lab.
To demonstrate their cell-free platform technology, the researchers produced a human kinase that is involved in tumor cell proliferation and showed that it was functional and active. Kinase is an enzyme (a protein acting as a catalyst) that transfers a phosphate group onto a protein. Through this process, kinases activate the function of proteins within the cell. Kinases are implicated in many diseases and, therefore, of particular interest.
“The ability to produce kinases for study should be useful in learning how these proteins function and in developing new types of drugs,” Jewett said.
Abstract
Understanding the functional and structural consequences of site-specific protein phosphorylation has remained limited by our inability to produce phosphoproteins at high yields. Here we address this limitation by developing a cell-free protein synthesis (CFPS) platform that employs crude extracts from a genomically recoded strain of Escherichia coli for site-specific, co-translational incorporation of phosphoserine into proteins. We apply this system to the robust production of up to milligram quantities of human MEK1 kinase. Then, we recapitulate a physiological signalling cascade in vitro to evaluate the contributions of site-specific phosphorylation of mono- and doubly phosphorylated forms on MEK1 activity. We discover that only one phosphorylation event is necessary and sufficient for MEK1 activity. Our work sets the stage for using CFPS as a rapid high-throughput technology platform for direct expression of programmable phosphoproteins containing multiple phosphorylated residues. This work will facilitate study of phosphorylation-dependent structure–function relationships, kinase signalling networks and kinase inhibitor drugs.
SOURCES – Northwestern University, Nature Communication

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.