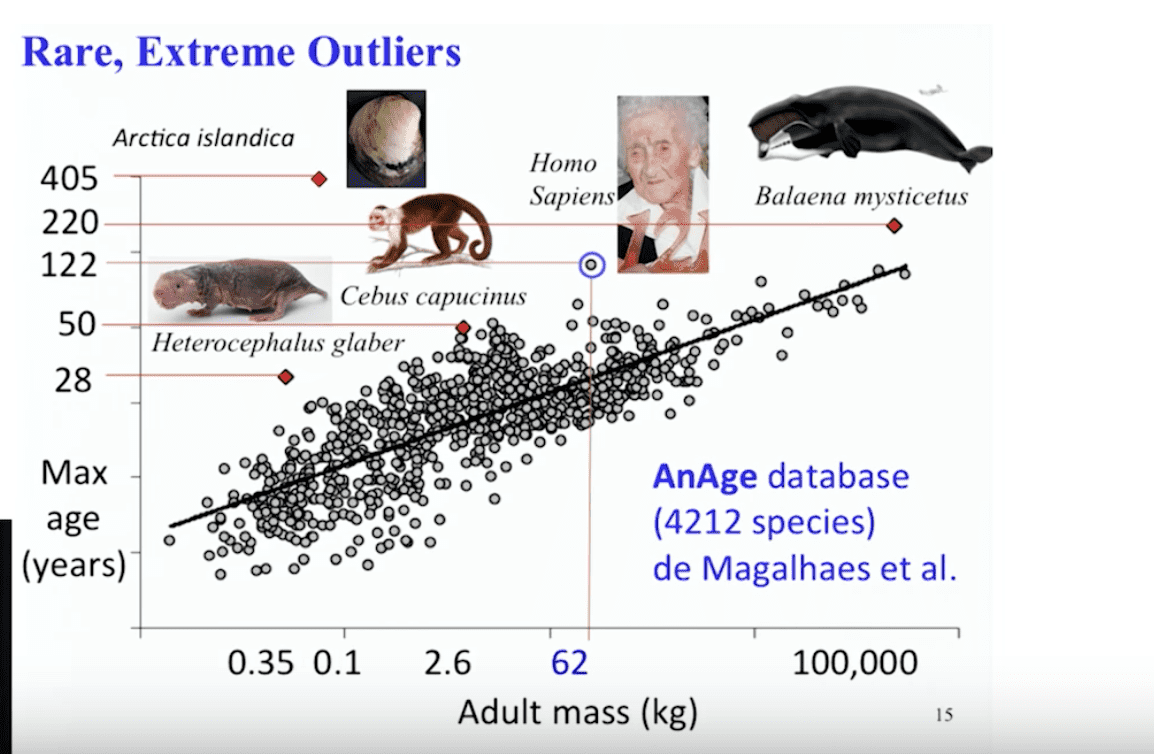
AnAge is a curated database of aging and life history in animals, including extensive longevity records. AnAge was primarily developed for comparative biology studies, in particular studies of longevity and aging, but can also be useful for ecological and conservation studies and as a reference for zoos and field biologists.
It can be used to find outlier animals with unusual longevity.
The ocean quahog (Arctica islandica) is a species of edible clam. Arctica islandica resembles the quahog, but the shell of the ocean quahog is rounder, the periostracum is usually black, and on the interior of the shell, the pallial line has no indentation, or sinus. Unlike the quahog, which lives intertidally and can be collected by clam digging, this species lives subtidally, and can only be collected by dredging. They grow to sizes exceeding 50 mm shell height. An individual specimen was reported to have lived 507 years.

Certain Koi fish have lived to 226 years of age.

Adwaita, an Aldabra giant tortoise, died at an estimated age of 255 in March 2006 in Alipore Zoo, Kolkata, India. If verified, it will have been the oldest terrestrial animal in the world.
Currently there are 4,244 entries (4,219 species and 25 taxa).
There are 3776 animal longevity records in the database.
Seven species of animals that do not seem to age

Longevity Map
LongevityMap is a database of human genetic variants associated with longevity. Negative results are also included in the LongevityMap to provide visitors with as much information as possible regarding each gene and variant previously studied in context of longevity. As such, the LongevityMap serves as a repository of genetic association studies of longevity and reflects our current knowledge of the genetics of human longevity.
LongevityMap can be done by gene or genetic variant.
There are currently 274 significant aging related genes.
Dietary restriction related genes
GenDR Database of Dietary Restriction-Related Genes
Dietary restriction (DR), limiting nutrient intake from diet without causing malnutrition, retards age-related degeneration and extends lifespan in multiple organisms. DR induces multiple changes, yet its underlying mechanisms remain poorly understood. To facilitate research on the genetic and molecular mechanisms of DR-induced life-extension, we developed GenDR, a database of genes associated with DR. GenDR includes two datasets: 1) genes inferred from experiments in model organisms in which genetic manipulations cancel out or disrupt the life-extending effects of DR; 2) genes robustly altered due to DR, derived from a meta-analysis of microarray DR studies in mammals. An analysis of the gene network of DR has also been performed using GenDR.
Understanding the genetic basis of DR is of great importance not only to the biology of ageing but to understand how diet can influence aging, longevity, health and age-related diseases. Pharmaceutical interventions that target DR-associated genes are also an emerging area with huge potential.
GenAge Database of Ageing-Related Genes
GenAge is the benchmark database of genes related to aging. GenAge is divided into genes related to longevity and/or aging in model organisms (yeast, worms, flies, mice, etc.) and aging-related human genes. The section on human aging-related genes includes the few genes directly related to aging in humans plus the best candidate genes obtained from model organisms. Human genes are thus considerably better annotated and include more information. GenAge is manually curated by experts to ensure high-quality content.
DrugAge Database of Anti-Aging Drugs
The DrugAge database contains an extensive compilation of drugs, compounds and supplements (including natural products and nutraceuticals) with anti-aging properties that extend longevity in model organisms. Our focus is on drugs/compounds potentially impacting on aging, and therefore drugs/compounds extending lifespan in disease-prone animals (e.g., cancer models) are excluded.
Lifespan increasing assays: 1,035
Lifespan decreasing assays: 201
Distinct drugs: 418
Distinct species: 27
CellAge: The Database of Cell Senescence Genes
Cell senescence can be defined as the irreversible cessation of cell division of normally proliferating cells. Human cells become senescent from progressive shortening of telomeres as cells divide, stress or oncogenes. Primarily an anti-tumour mechanism, senescent cells accumulate with age in tissues and have been associated with degeneration and ageing of whole organisms. Many proteins have been linked to cell senescence as biomarkers and as causal drivers. To facilitate studies focused on cell senescence, we developed CellAge, a database of genes associated with cell senescence. Our manually-curated data is based on gene manipulation experiments in different human cell types. A gene expression signature of cellular senescence will also be made available in due course.
CellAge is in the beta phase of development and therefore still being improved and expanded.

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.

Dont worry. They have 2 turns max. Basically for that reason.
wiki plagiarism describing the clam
The Great Basin bristlecone pine is an animal?
Are we really sure the extending the life of, say, Donald Trump is a good thing ? Really really sure?
Dont worry. They have 2 turns max. Basically for that reason.
wiki plagiarism describing the clam
The Great Basin bristlecone pine is an animal?
Are we really sure the extending the life of say Donald Trump is a good thing ? Really really sure?
Trump could step down for a term and then be re-elected for a third and fourth term – he just needs to step aside for one term. I say, make Trump POTUS for life.
Trump could step down for a term and then be re-elected for a third and fourth term – he just needs to step aside for one term. I say make Trump POTUS for life.
I still think you should have emojis for “hopeful” and “scared out of my wits”…
Maybe they’ll change the law in the second turn. 😛
Turtles and fish are not that far.
Seven species of animals that do not seem to age” Also all single celled organisms.
The naked mole rats have already demonstrated some interesting biological features, such as a modified ribosome with a much lower error rate. However, I wouldn’t count them as warm blooded. They’re one of the rare cold blooded mammals.
Hm, I think that studying large long-lived mammals (Bowhead whale), and other exceptionally long-lived warm-blooded animals (parrots!, naked mole-rats) would be a better pathway than trees and various more ‘distant’ animals. Apparently these animals have found a way to mitigate senescence and genetic deterioration (cancer), while having a fast warm-blooded metabolism.
It’s ok, it isn’t immortality. It is life extension. Accidental death and murder will still work in the future.
I still think you should have emojis for hopeful”” and “”””scared out of my wits””””…”””
Maybe they’ll change the law in the second turn. 😛
Turtles and fish are not that far.
Seven species of animals that do not seem to age””Also all single celled organisms.”””
The naked mole rats have already demonstrated some interesting biological features such as a modified ribosome with a much lower error rate.However I wouldn’t count them as warm blooded. They’re one of the rare cold blooded mammals.
Hm I think that studying large long-lived mammals (Bowhead whale) and other exceptionally long-lived warm-blooded animals (parrots! naked mole-rats) would be a better pathway than trees and various more ‘distant’ animals.Apparently these animals have found a way to mitigate senescence and genetic deterioration (cancer) while having a fast warm-blooded metabolism.
It’s ok it isn’t immortality. It is life extension. Accidental death and murder will still work in the future.
Would that be enough for the second civil war they are fearing right now?
Would that be enough for the second civil war they are fearing right now?
Really? They are very distant genetically, and the long-lived ones tend to live in cold regions with poor food supplies. That’s not going to help us out as much as actual mammals, with similar blood types, air breathing, all that.
And skeptical. Indifferent is close, I suppose. And the picture that goes with excited is laughing, so I think of it (right or wrong) as laughing at the concepts proposed.
Civil War. Until Americans have burgers to stuff their faces with there won’t be any civil war.
Really? They are very distant genetically and the long-lived ones tend to live in cold regions with poor food supplies. That’s not going to help us out as much as actual mammals with similar blood types air breathing all that.
And skeptical. Indifferent is close I suppose. And the picture that goes with excited is laughing so I think of it (right or wrong) as laughing at the concepts proposed.
Civil War. Until Americans have burgers to stuff their faces with there won’t be any civil war.
How would family relations work when your great-grandmother looks as young as you? ” (and ultra hot)
Imagine a world where humans do not age. How would family relations work when your great-grandmother looks as young as you? How family ties would develop among multiple generations, all of which seem to be the same age? Would inheritance be a thing of the past? Would patricide (or matricide) increase, as progeny waits 200 years to take control of that megacorp, but the unaging CEO, the patriarch or matriarch, is in good health and looking like he/she is in his 30s?
LOL, good point. Also, the “animals” that don’t seem to age includes animals that don’t live past the century mark, so I’m not sure how they can make that claim. Just being long-lived is not the same as not aging. And if you’re going to put turtles on the list, I would start not with Blandings (even a snapper lives longer), but with Galapagos Tortoise, or the Aldabra giant tortoises, which can apparently live over 200 years. Meanwhile, they apparently totally ignored the “immortal jellyfish’ (Turritopsis dohrnii) that can revert back to its infant state and restart its life cycle. IMO, the Wikipedia pages on “biological immortality” and “list of longest-living organisms” are much better than this group’s spreadsheet on animals. Hopefully their genetic database has more thought behind it.
How would family relations work when your great-grandmother looks as young as you? “” (and ultra hot)”””
Imagine a world where humans do not age. How would family relations work when your great-grandmother looks as young as you?How family ties would develop among multiple generations all of which seem to be the same age?Would inheritance be a thing of the past? Would patricide (or matricide) increase as progeny waits 200 years to take control of that megacorp but the unaging CEO the patriarch or matriarch is in good health and looking like he/she is in his 30s?
LOL good point. Also the animals”” that don’t seem to age includes animals that don’t live past the century mark”” so I’m not sure how they can make that claim. Just being long-lived is not the same as not aging. And if you’re going to put turtles on the list I would start not with Blandings (even a snapper lives longer) but with Galapagos Tortoise or the Aldabra giant tortoises which can apparently live over 200 years. Meanwhile”” they apparently totally ignored the “”””immortal jellyfish’ (Turritopsis dohrnii) that can revert back to its infant state and restart its life cycle.IMO”””” the Wikipedia pages on “”””biological immortality”””” and “”””list of longest-living organisms”””” are much better than this group’s spreadsheet on animals. Hopefully their genetic database has more thought behind it.”””
Fearing” ? Everyone I’ve seen writing about the prospect seems to look forward to it. Because everyone thinks they are secretly in the vast majority. And that everyone on the other side is a combination of completely incompetent and a hopeless coward who will turn and flee back to their trailer park/ghetto/university/inner-city-apartment/suburban house/housing project/back-across-the-border whichever is appropriate.
MAGA!! Trump 2020!!
Fearing”” ?Everyone I’ve seen writing about the prospect seems to look forward to it. Because everyone thinks they are secretly in the vast majority. And that everyone on the other side is a combination of completely incompetent and a hopeless coward who will turn and flee back to their trailer park/ghetto/university/inner-city-apartment/suburban house/housing project/back-across-the-border whichever is appropriate.”””
MAGA!! Trump 2020!!
It would be impossible to allow for everyone, or the result would be massive overpopulation. I imagine, in USA you would buy the right to have a child on an auction, put up whenever some dies accidentally, or takes off into space. In Europe it would be a lottery.
It would be impossible to allow for everyone or the result would be massive overpopulation.I imagine in USA you would buy the right to have a child on an auction put up whenever some dies accidentally or takes off into space.In Europe it would be a lottery.
If you run the numbers, with current global death and birth rates, it will take ~66 years to double the population. If all death are eliminated tomorrow (same birth rate, 0 death rate; not going to happen even with anti-aging), then it would take 38 years to double, IIRC. That not a huge difference. Then you need to take into account that birth rates are declining. With a much longer breeding age range, it will likely decline even more, so the difference will be even smaller. Your overpopulation panic is wildly exaggerated.
If you run the numbers with current global death and birth rates it will take ~66 years to double the population. If all death are eliminated tomorrow (same birth rate 0 death rate; not going to happen even with anti-aging) then it would take 38 years to double IIRC. That not a huge difference.Then you need to take into account that birth rates are declining. With a much longer breeding age range it will likely decline even more so the difference will be even smaller.Your overpopulation panic is wildly exaggerated.
It won’t flatten out (probably), but it can come pretty close. Regardless, the point is that population growth takes a long time, during which society can adapt in all sorts of ways. It’s not going to explode overnight. I didn’t want to go into an analysis of the available resources etc, to avoid distracting from that basic math, but there are enough resources even down here on Earth for possibly as many as 100 billion, if not more. The key is correct *management*, reducing pollution, recycling, etc. We’re gradually getting better at that. Specifically to your question on crowding, at 10000 people/km^2 – slightly less than NYC population density – 100 billion would take 10 million km^2. That’s ~7% of Earth’s land area. A bit much, perhaps, but hardly catastrophic. If we adopt seasteading, then it’s ~3% of the ocean area, or ~2% of Earth’s total surface. We can also build down and up to add more space. I personally like the idea of 3D cities (similar to the concept of “archologies” from SimCity). But as you say, we’ll be settling space in the coming decades. There’s orders of magnitude more room and resources there. One should also keep in mind that it’ll likely take a few decades before anti-aging can be widely deployed – which gives extra time for birth rates to decline, for environmental controls to improve, for space technologies to mature, etc. Btw, here some more math: Assuming we start from 8 billion a bit in the future, getting to 100 billion would take about 3.5 doublings. At the worst case of ~40 years per doubling (today’s birth rate, 0 death rate), it would take ~140 years. At 50 years per doubling (slightly lower birth rate), it would take 175 years. So we won’t hit those numbers until ~2200 or later. That’s plenty of time to settle space, as well as fix things down here. But the doubling rate could be even lower than, and even one doubling of 40 years is a pretty long time.
Current population projections tend to top out at about 12 billion people sometime in the 22 century, but with immortality, this curve will never flatten out, but keep rising. How many people would you like in your city? Your timeframe is too short to see the issue. Then again, my timeframe opens up future tech. After all, there is plenty of space in the belt, so i guess its not a problem.
It won’t flatten out (probably) but it can come pretty close. Regardless the point is that population growth takes a long time during which society can adapt in all sorts of ways. It’s not going to explode overnight.I didn’t want to go into an analysis of the available resources etc to avoid distracting from that basic math but there are enough resources even down here on Earth for possibly as many as 100 billion if not more. The key is correct *management* reducing pollution recycling etc. We’re gradually getting better at that.Specifically to your question on crowding at 10000 people/km^2 – slightly less than NYC population density – 100 billion would take 10 million km^2. That’s ~7{22800fc54956079738b58e74e4dcd846757aa319aad70fcf90c97a58f3119a12} of Earth’s land area. A bit much perhaps but hardly catastrophic. If we adopt seasteading then it’s ~3{22800fc54956079738b58e74e4dcd846757aa319aad70fcf90c97a58f3119a12} of the ocean area or ~2{22800fc54956079738b58e74e4dcd846757aa319aad70fcf90c97a58f3119a12} of Earth’s total surface. We can also build down and up to add more space. I personally like the idea of 3D cities (similar to the concept of archologies”” from SimCity).But as you say”” we’ll be settling space in the coming decades. There’s orders of magnitude more room and resources there.One should also keep in mind that it’ll likely take a few decades before anti-aging can be widely deployed – which gives extra time for birth rates to decline for environmental controls to improve for space technologies to mature etc.Btw here some more math:Assuming we start from 8 billion a bit in the future getting to 100 billion would take about 3.5 doublings. At the worst case of ~40 years per doubling (today’s birth rate 0 death rate) it would take ~140 years. At 50 years per doubling (slightly lower birth rate) it would take 175 years. So we won’t hit those numbers until ~2200 or later. That’s plenty of time to settle space as well as”
Current population projections tend to top out at about 12 billion people sometime in the 22 century but with immortality this curve will never flatten out but keep rising. How many people would you like in your city?Your timeframe is too short to see the issue.Then again my timeframe opens up future tech. After all there is plenty of space in the belt so i guess its not a problem.
Based on income, you are a global one percenter if you make more that $32,400 a year. More than half the world does not have $2,300 in wealth. Even if unlimited life extension were to cost no more than a caffeinated beverage at Starbucks each day, It might be out of reach of the vast majority of the world. Then too, I know people who wouldn’t do it just because it might involve taking a pill every day and they would either forget or tell themselves, “Well, skipping one day won’t matter.” And just imagine if it required going to the gym for 45 minutes every day! Among those who do get this life extension, half life of the human population would be about 1,000 years, providing accidental deaths and murders don’t change substantially. Some would probably lock themselves in underground bunkers and operate through surrogate bodies (when that technology matures) to reduce their chances of accidents. Finally, there would be very little turnover at the top. Regardless of their personal income or wealth, most ambitious folks, people who want to take charge and run things, would almost inevitably have to emigrate off planet, perhaps even out of the solar system, if that ever becomes a thing.
Based on income you are a global one percenter if you make more that $32400 a year. More than half the world does not have $2300 in wealth. Even if unlimited life extension were to cost no more than a caffeinated beverage at Starbucks each day It might be out of reach of the vast majority of the world. Then too I know people who wouldn’t do it just because it might involve taking a pill every day and they would either forget or tell themselves Well”” skipping one day won’t matter.”” And just imagine if it required going to the gym for 45 minutes every day!Among those who do get this life extension”” half life of the human population would be about 1000 years providing accidental deaths and murders don’t change substantially. Some would probably lock themselves in underground bunkers and operate through surrogate bodies (when that technology matures) to reduce their chances of accidents.Finally there would be very little turnover at the top. Regardless of their personal income or wealth most ambitious folks people who want to take charge and run things would almost inevitably have to emigrate off planet perhaps even out of the solar system”” if that ever becomes a thing.”””
Based on income, you are a global one percenter if you make more that $32,400 a year. More than half the world does not have $2,300 in wealth. Even if unlimited life extension were to cost no more than a caffeinated beverage at Starbucks each day, It might be out of reach of the vast majority of the world. Then too, I know people who wouldn’t do it just because it might involve taking a pill every day and they would either forget or tell themselves, “Well, skipping one day won’t matter.” And just imagine if it required going to the gym for 45 minutes every day! Among those who do get this life extension, half life of the human population would be about 1,000 years, providing accidental deaths and murders don’t change substantially. Some would probably lock themselves in underground bunkers and operate through surrogate bodies (when that technology matures) to reduce their chances of accidents. Finally, there would be very little turnover at the top. Regardless of their personal income or wealth, most ambitious folks, people who want to take charge and run things, would almost inevitably have to emigrate off planet, perhaps even out of the solar system, if that ever becomes a thing.
Based on income you are a global one percenter if you make more that $32400 a year. More than half the world does not have $2300 in wealth. Even if unlimited life extension were to cost no more than a caffeinated beverage at Starbucks each day It might be out of reach of the vast majority of the world. Then too I know people who wouldn’t do it just because it might involve taking a pill every day and they would either forget or tell themselves Well”” skipping one day won’t matter.”” And just imagine if it required going to the gym for 45 minutes every day!Among those who do get this life extension”” half life of the human population would be about 1000 years providing accidental deaths and murders don’t change substantially. Some would probably lock themselves in underground bunkers and operate through surrogate bodies (when that technology matures) to reduce their chances of accidents.Finally there would be very little turnover at the top. Regardless of their personal income or wealth most ambitious folks people who want to take charge and run things would almost inevitably have to emigrate off planet perhaps even out of the solar system”” if that ever becomes a thing.”””
Based on income, you are a global one percenter if you make more that $32,400 a year. More than half the world does not have $2,300 in wealth. Even if unlimited life extension were to cost no more than a caffeinated beverage at Starbucks each day, It might be out of reach of the vast majority of the world. Then too, I know people who wouldn’t do it just because it might involve taking a pill every day and they would either forget or tell themselves, “Well, skipping one day won’t matter.” And just imagine if it required going to the gym for 45 minutes every day!
Among those who do get this life extension, half life of the human population would be about 1,000 years, providing accidental deaths and murders don’t change substantially. Some would probably lock themselves in underground bunkers and operate through surrogate bodies (when that technology matures) to reduce their chances of accidents.
Finally, there would be very little turnover at the top. Regardless of their personal income or wealth, most ambitious folks, people who want to take charge and run things, would almost inevitably have to emigrate off planet, perhaps even out of the solar system, if that ever becomes a thing.
It won’t flatten out (probably), but it can come pretty close. Regardless, the point is that population growth takes a long time, during which society can adapt in all sorts of ways. It’s not going to explode overnight. I didn’t want to go into an analysis of the available resources etc, to avoid distracting from that basic math, but there are enough resources even down here on Earth for possibly as many as 100 billion, if not more. The key is correct *management*, reducing pollution, recycling, etc. We’re gradually getting better at that. Specifically to your question on crowding, at 10000 people/km^2 – slightly less than NYC population density – 100 billion would take 10 million km^2. That’s ~7% of Earth’s land area. A bit much, perhaps, but hardly catastrophic. If we adopt seasteading, then it’s ~3% of the ocean area, or ~2% of Earth’s total surface. We can also build down and up to add more space. I personally like the idea of 3D cities (similar to the concept of “archologies” from SimCity). But as you say, we’ll be settling space in the coming decades. There’s orders of magnitude more room and resources there. One should also keep in mind that it’ll likely take a few decades before anti-aging can be widely deployed – which gives extra time for birth rates to decline, for environmental controls to improve, for space technologies to mature, etc. Btw, here some more math: Assuming we start from 8 billion a bit in the future, getting to 100 billion would take about 3.5 doublings. At the worst case of ~40 years per doubling (today’s birth rate, 0 death rate), it would take ~140 years. At 50 years per doubling (slightly lower birth rate), it would take 175 years. So we won’t hit those numbers until ~2200 or later. That’s plenty of time to settle space, as well as fix things down here. But the doubling rate could be even lower than, and even one doubling of 40 years is a pretty long time.
It won’t flatten out (probably) but it can come pretty close. Regardless the point is that population growth takes a long time during which society can adapt in all sorts of ways. It’s not going to explode overnight.I didn’t want to go into an analysis of the available resources etc to avoid distracting from that basic math but there are enough resources even down here on Earth for possibly as many as 100 billion if not more. The key is correct *management* reducing pollution recycling etc. We’re gradually getting better at that.Specifically to your question on crowding at 10000 people/km^2 – slightly less than NYC population density – 100 billion would take 10 million km^2. That’s ~7{22800fc54956079738b58e74e4dcd846757aa319aad70fcf90c97a58f3119a12} of Earth’s land area. A bit much perhaps but hardly catastrophic. If we adopt seasteading then it’s ~3{22800fc54956079738b58e74e4dcd846757aa319aad70fcf90c97a58f3119a12} of the ocean area or ~2{22800fc54956079738b58e74e4dcd846757aa319aad70fcf90c97a58f3119a12} of Earth’s total surface. We can also build down and up to add more space. I personally like the idea of 3D cities (similar to the concept of archologies”” from SimCity).But as you say”” we’ll be settling space in the coming decades. There’s orders of magnitude more room and resources there.One should also keep in mind that it’ll likely take a few decades before anti-aging can be widely deployed – which gives extra time for birth rates to decline for environmental controls to improve for space technologies to mature etc.Btw here some more math:Assuming we start from 8 billion a bit in the future getting to 100 billion would take about 3.5 doublings. At the worst case of ~40 years per doubling (today’s birth rate 0 death rate) it would take ~140 years. At 50 years per doubling (slightly lower birth rate) it would take 175 years. So we won’t hit those numbers until ~2200 or later. That’s plenty of time to settle space as well as”
Current population projections tend to top out at about 12 billion people sometime in the 22 century, but with immortality, this curve will never flatten out, but keep rising. How many people would you like in your city? Your timeframe is too short to see the issue. Then again, my timeframe opens up future tech. After all, there is plenty of space in the belt, so i guess its not a problem.
Current population projections tend to top out at about 12 billion people sometime in the 22 century but with immortality this curve will never flatten out but keep rising. How many people would you like in your city?Your timeframe is too short to see the issue.Then again my timeframe opens up future tech. After all there is plenty of space in the belt so i guess its not a problem.
It won’t flatten out (probably), but it can come pretty close. Regardless, the point is that population growth takes a long time, during which society can adapt in all sorts of ways. It’s not going to explode overnight.
I didn’t want to go into an analysis of the available resources etc, to avoid distracting from that basic math, but there are enough resources even down here on Earth for possibly as many as 100 billion, if not more. The key is correct *management*, reducing pollution, recycling, etc. We’re gradually getting better at that.
Specifically to your question on crowding, at 10000 people/km^2 – slightly less than NYC population density – 100 billion would take 10 million km^2. That’s ~7% of Earth’s land area. A bit much, perhaps, but hardly catastrophic. If we adopt seasteading, then it’s ~3% of the ocean area, or ~2% of Earth’s total surface. We can also build down and up to add more space. I personally like the idea of 3D cities (similar to the concept of “archologies” from SimCity).
But as you say, we’ll be settling space in the coming decades. There’s orders of magnitude more room and resources there.
One should also keep in mind that it’ll likely take a few decades before anti-aging can be widely deployed – which gives extra time for birth rates to decline, for environmental controls to improve, for space technologies to mature, etc.
Btw, here some more math:
Assuming we start from 8 billion a bit in the future, getting to 100 billion would take about 3.5 doublings. At the worst case of ~40 years per doubling (today’s birth rate, 0 death rate), it would take ~140 years. At 50 years per doubling (slightly lower birth rate), it would take 175 years. So we won’t hit those numbers until ~2200 or later. That’s plenty of time to settle space, as well as fix things down here. But the doubling rate could be even lower than, and even one doubling of 40 years is a pretty long time.
Current population projections tend to top out at about 12 billion people sometime in the 22 century, but with immortality, this curve will never flatten out, but keep rising. How many people would you like in your city?
Your timeframe is too short to see the issue.
Then again, my timeframe opens up future tech. After all, there is plenty of space in the belt, so i guess its not a problem.
If you run the numbers, with current global death and birth rates, it will take ~66 years to double the population. If all death are eliminated tomorrow (same birth rate, 0 death rate; not going to happen even with anti-aging), then it would take 38 years to double, IIRC. That not a huge difference. Then you need to take into account that birth rates are declining. With a much longer breeding age range, it will likely decline even more, so the difference will be even smaller. Your overpopulation panic is wildly exaggerated.
If you run the numbers with current global death and birth rates it will take ~66 years to double the population. If all death are eliminated tomorrow (same birth rate 0 death rate; not going to happen even with anti-aging) then it would take 38 years to double IIRC. That not a huge difference.Then you need to take into account that birth rates are declining. With a much longer breeding age range it will likely decline even more so the difference will be even smaller.Your overpopulation panic is wildly exaggerated.
If you run the numbers, with current global death and birth rates, it will take ~66 years to double the population. If all death are eliminated tomorrow (same birth rate, 0 death rate; not going to happen even with anti-aging), then it would take 38 years to double, IIRC. That not a huge difference.
Then you need to take into account that birth rates are declining. With a much longer breeding age range, it will likely decline even more, so the difference will be even smaller.
Your overpopulation panic is wildly exaggerated.
It would be impossible to allow for everyone, or the result would be massive overpopulation. I imagine, in USA you would buy the right to have a child on an auction, put up whenever some dies accidentally, or takes off into space. In Europe it would be a lottery.
It would be impossible to allow for everyone or the result would be massive overpopulation.I imagine in USA you would buy the right to have a child on an auction put up whenever some dies accidentally or takes off into space.In Europe it would be a lottery.
Fearing” ? Everyone I’ve seen writing about the prospect seems to look forward to it. Because everyone thinks they are secretly in the vast majority. And that everyone on the other side is a combination of completely incompetent and a hopeless coward who will turn and flee back to their trailer park/ghetto/university/inner-city-apartment/suburban house/housing project/back-across-the-border whichever is appropriate.
Fearing”” ?Everyone I’ve seen writing about the prospect seems to look forward to it. Because everyone thinks they are secretly in the vast majority. And that everyone on the other side is a combination of completely incompetent and a hopeless coward who will turn and flee back to their trailer park/ghetto/university/inner-city-apartment/suburban house/housing project/back-across-the-border whichever is appropriate.”””
MAGA!! Trump 2020!!
MAGA!! Trump 2020!!
It would be impossible to allow for everyone, or the result would be massive overpopulation.
I imagine, in USA you would buy the right to have a child on an auction, put up whenever some dies accidentally, or takes off into space.
In Europe it would be a lottery.
How would family relations work when your great-grandmother looks as young as you? ” (and ultra hot)
How would family relations work when your great-grandmother looks as young as you? “” (and ultra hot)”””
Imagine a world where humans do not age. How would family relations work when your great-grandmother looks as young as you? How family ties would develop among multiple generations, all of which seem to be the same age? Would inheritance be a thing of the past? Would patricide (or matricide) increase, as progeny waits 200 years to take control of that megacorp, but the unaging CEO, the patriarch or matriarch, is in good health and looking like he/she is in his 30s?
Imagine a world where humans do not age. How would family relations work when your great-grandmother looks as young as you?How family ties would develop among multiple generations all of which seem to be the same age?Would inheritance be a thing of the past? Would patricide (or matricide) increase as progeny waits 200 years to take control of that megacorp but the unaging CEO the patriarch or matriarch is in good health and looking like he/she is in his 30s?
LOL, good point. Also, the “animals” that don’t seem to age includes animals that don’t live past the century mark, so I’m not sure how they can make that claim. Just being long-lived is not the same as not aging. And if you’re going to put turtles on the list, I would start not with Blandings (even a snapper lives longer), but with Galapagos Tortoise, or the Aldabra giant tortoises, which can apparently live over 200 years. Meanwhile, they apparently totally ignored the “immortal jellyfish’ (Turritopsis dohrnii) that can revert back to its infant state and restart its life cycle. IMO, the Wikipedia pages on “biological immortality” and “list of longest-living organisms” are much better than this group’s spreadsheet on animals. Hopefully their genetic database has more thought behind it.
LOL good point. Also the animals”” that don’t seem to age includes animals that don’t live past the century mark”” so I’m not sure how they can make that claim. Just being long-lived is not the same as not aging. And if you’re going to put turtles on the list I would start not with Blandings (even a snapper lives longer) but with Galapagos Tortoise or the Aldabra giant tortoises which can apparently live over 200 years. Meanwhile”” they apparently totally ignored the “”””immortal jellyfish’ (Turritopsis dohrnii) that can revert back to its infant state and restart its life cycle.IMO”””” the Wikipedia pages on “”””biological immortality”””” and “”””list of longest-living organisms”””” are much better than this group’s spreadsheet on animals. Hopefully their genetic database has more thought behind it.”””
Really? They are very distant genetically, and the long-lived ones tend to live in cold regions with poor food supplies. That’s not going to help us out as much as actual mammals, with similar blood types, air breathing, all that.
Really? They are very distant genetically and the long-lived ones tend to live in cold regions with poor food supplies. That’s not going to help us out as much as actual mammals with similar blood types air breathing all that.
And skeptical. Indifferent is close, I suppose. And the picture that goes with excited is laughing, so I think of it (right or wrong) as laughing at the concepts proposed.
And skeptical. Indifferent is close I suppose. And the picture that goes with excited is laughing so I think of it (right or wrong) as laughing at the concepts proposed.
Civil War. Until Americans have burgers to stuff their faces with there won’t be any civil war.
Civil War. Until Americans have burgers to stuff their faces with there won’t be any civil war.
Would that be enough for the second civil war they are fearing right now?
Would that be enough for the second civil war they are fearing right now?
I still think you should have emojis for “hopeful” and “scared out of my wits”…
I still think you should have emojis for hopeful”” and “”””scared out of my wits””””…”””
Maybe they’ll change the law in the second turn. 😛
Maybe they’ll change the law in the second turn. 😛
Turtles and fish are not that far.
Turtles and fish are not that far.
Seven species of animals that do not seem to age” Also all single celled organisms.
Seven species of animals that do not seem to age””Also all single celled organisms.”””
“Fearing” ?
Everyone I’ve seen writing about the prospect seems to look forward to it. Because everyone thinks they are secretly in the vast majority. And that everyone on the other side is a combination of completely incompetent and a hopeless coward who will turn and flee back to their trailer park/ghetto/university/inner-city-apartment/suburban house/housing project/back-across-the-border whichever is appropriate.
The naked mole rats have already demonstrated some interesting biological features, such as a modified ribosome with a much lower error rate. However, I wouldn’t count them as warm blooded. They’re one of the rare cold blooded mammals.
The naked mole rats have already demonstrated some interesting biological features such as a modified ribosome with a much lower error rate.However I wouldn’t count them as warm blooded. They’re one of the rare cold blooded mammals.
MAGA!! Trump 2020!!
Hm, I think that studying large long-lived mammals (Bowhead whale), and other exceptionally long-lived warm-blooded animals (parrots!, naked mole-rats) would be a better pathway than trees and various more ‘distant’ animals. Apparently these animals have found a way to mitigate senescence and genetic deterioration (cancer), while having a fast warm-blooded metabolism.
Hm I think that studying large long-lived mammals (Bowhead whale) and other exceptionally long-lived warm-blooded animals (parrots! naked mole-rats) would be a better pathway than trees and various more ‘distant’ animals.Apparently these animals have found a way to mitigate senescence and genetic deterioration (cancer) while having a fast warm-blooded metabolism.
It’s ok, it isn’t immortality. It is life extension. Accidental death and murder will still work in the future.
It’s ok it isn’t immortality. It is life extension. Accidental death and murder will still work in the future.
Trump could step down for a term and then be re-elected for a third and fourth term – he just needs to step aside for one term. I say, make Trump POTUS for life.
Trump could step down for a term and then be re-elected for a third and fourth term – he just needs to step aside for one term. I say make Trump POTUS for life.
Dont worry. They have 2 turns max. Basically for that reason.
Dont worry. They have 2 turns max. Basically for that reason.
wiki plagiarism describing the clam
wiki plagiarism describing the clam
The Great Basin bristlecone pine is an animal?
The Great Basin bristlecone pine is an animal?
“How would family relations work when your great-grandmother looks as young as you? ” (and ultra hot)
Imagine a world where humans do not age.
How would family relations work when your great-grandmother looks as young as you?
How family ties would develop among multiple generations, all of which seem to be the same age?
Would inheritance be a thing of the past? Would patricide (or matricide) increase, as progeny waits 200 years to take control of that megacorp, but the unaging CEO, the patriarch or matriarch, is in good health and looking like he/she is in his 30s?
LOL, good point. Also, the “animals” that don’t seem to age includes animals that don’t live past the century mark, so I’m not sure how they can make that claim. Just being long-lived is not the same as not aging. And if you’re going to put turtles on the list, I would start not with Blandings (even a snapper lives longer), but with Galapagos Tortoise, or the Aldabra giant tortoises, which can apparently live over 200 years.
Meanwhile, they apparently totally ignored the “immortal jellyfish’ (Turritopsis dohrnii) that can revert back to its infant state and restart its life cycle.
IMO, the Wikipedia pages on “biological immortality” and “list of longest-living organisms” are much better than this group’s spreadsheet on animals. Hopefully their genetic database has more thought behind it.
Really? They are very distant genetically, and the long-lived ones tend to live in cold regions with poor food supplies. That’s not going to help us out as much as actual mammals, with similar blood types, air breathing, all that.
And skeptical. Indifferent is close, I suppose. And the picture that goes with excited is laughing, so I think of it (right or wrong) as laughing at the concepts proposed.
Civil War. Until Americans have burgers to stuff their faces with there won’t be any civil war.
Are we really sure the extending the life of, say, Donald Trump is a good thing ? Really really sure?
Are we really sure the extending the life of say Donald Trump is a good thing ? Really really sure?
Would that be enough for the second civil war they are fearing right now?
I still think you should have emojis for “hopeful” and “scared out of my wits”…
Maybe they’ll change the law in the second turn. 😛
Turtles and fish are not that far.
“Seven species of animals that do not seem to age”
Also all single celled organisms.
The naked mole rats have already demonstrated some interesting biological features, such as a modified ribosome with a much lower error rate.
However, I wouldn’t count them as warm blooded. They’re one of the rare cold blooded mammals.
Hm, I think that studying large long-lived mammals (Bowhead whale), and other exceptionally long-lived warm-blooded animals (parrots!, naked mole-rats) would be a better pathway than trees and various more ‘distant’ animals.
Apparently these animals have found a way to mitigate senescence and genetic deterioration (cancer), while having a fast warm-blooded metabolism.
It’s ok, it isn’t immortality. It is life extension. Accidental death and murder will still work in the future.
Trump could step down for a term and then be re-elected for a third and fourth term – he just needs to step aside for one term. I say, make Trump POTUS for life.
Dont worry. They have 2 turns max. Basically for that reason.
wiki plagiarism describing the clam
The Great Basin bristlecone pine is an animal?
Are we really sure the extending the life of, say, Donald Trump is a good thing ?
Really really sure?