Chinese scientists used CRISPR gene therapy to correct a mutation that causes Marfan syndrome, an incurable connective tissue disorder that affects about 1 in 5,000 people. A single letter mistake in the gene for FBN1, which codes for the fibrillin protein, can cause a ripple effect of problems—from loose joints to weak vision to life-threatening tears in the heart’s walls. Starting with healthy eggs and sperm donated by a Marfan syndrome patient, the team of researchers from Shanghai Tech University and Guangzhou Medical University used an IVF technique to make viable human embryos. Then they injected the embryos with a Crispr construct known as a base editor, which swaps out a single DNA nucleotide for another—in this case, removing an “A” and replacing it with a “G”. They kept the embryos alive for another two days in the lab, long enough to run tests to see how well the editing worked.
Sequencing revealed that all 18 embryos had been edited, with 16 of the embryos bearing only the corrected version of the FBN1 gene. In two of the embryos, additional unwanted edits had also taken place. Previously, the most successful demonstration of gene editing in the human germline was the correction of a mutation that causes a hereditary heart condition in 42 out of 58 embryos. That study, which was published last year, used standard Crispr cut-and-paste technology.
Beam Technologies will be just one of many US companies looking at an increasingly streamlined path for genetic medicines. In July, FDA Commissioner Scott Gottlieb announced a new regulatory framework for gene therapies to treat rare diseases. The agency issued a suite of six guidance documents updating the approval process. And on August 17, the FDA along with the National Institutes of Health proposed changes in the way the agencies together assess the safety of gene-therapy human trials.
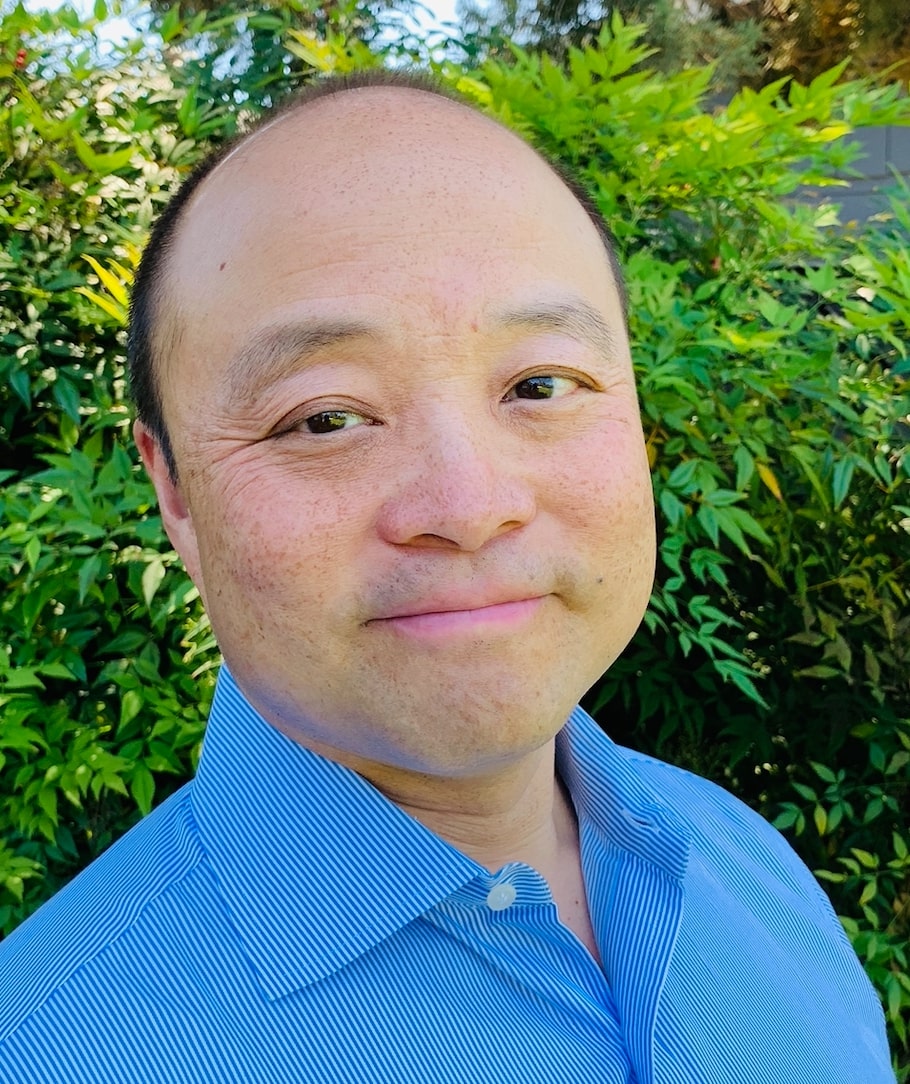
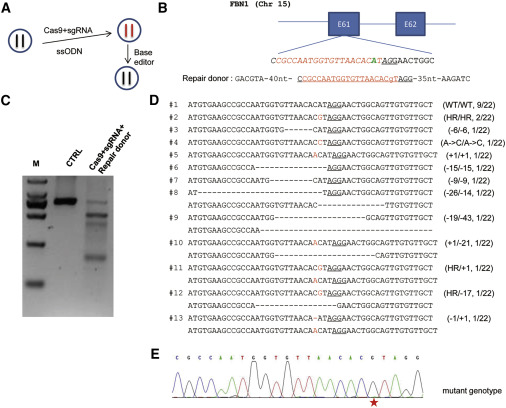
There are urgent demands for efficient treatment of heritable genetic diseases. The base editing technology has displayed its efficiency and precision in base substitution in human embryos, providing a potential early-stage treatment for genetic diseases. Taking advantage of this technology, we corrected a Marfan syndrome pathogenic mutation, FBN1T7498C. We first tested the feasibility in mutant cells, then successfully achieved genetic correction in heterozygous human embryos. The results showed that the BE3 mediated perfect correction at the efficiency of about 89%. Importantly, no off-target and indels were detected in any tested sites in samples by high-throughput deep sequencing combined with whole-genome sequencing analysis. Our study therefore suggests the efficiency and genetic safety of correcting a Marfan syndrome (MFS) pathogenic mutation in embryos by base editing.
Nearly 10,000 genetic diseases have been identified, which affects millions of families around the world. However, less than 6% of genetic diseases have approved treatments. To reduce the burden of genetic diseases in affected families and individuals, breakthroughs in diagnosis and therapeutic methods are urgently needed. Although preimplantation genetic diagnosis (PGD) is useful to prevent the generational transmission of the mutant allele to embryos, the potential risk for diagnostic errors still exists. On the other hand, gene therapy potentially provides an active approach for correcting the genetic diseases.
Genome editing technologies, especially those based on CRISRP/Cas9, have been successfully applied in genome manipulation, which has inspired a brilliant outlook that the pathogenic mutation can be precisely repaired to achieve therapeutic effects. Moreover, recent successes in precise genome editing trials in early human embryos have suggested a potentially true cure for genetic diseases. However, genome editing of human embryos caused huge concerns because of ethical issues and technical uncertainties regarding the efficiency and off-target effects. It is well-known that genome editing through CRISPR/Cas9 generates double-strand breaks (DSBs) that evoke the error-prone non-homologous end joining (NHEJ) DNA repair pathway after, which causes off-target mutagenesis. The recently developed base editor (BE) system was constructed by fusing the deaminase to the dCas9 protein. BE efficiently edits the mutation without donor, which is required for precise genome editing through homologous recombination at low efficiency. BE edits specific sites by C-to-T or G-to-A conversion without DSB formation, providing a safer genome editing tool with low off-target effects. BE has been applied in plants and animals, and has shown enormous advantages in precise base-level genome editing compared with CRISPR/Cas9 Based on the developments of the BE systems, several studies have been undertaken and have proven the efficiency and safety of BE in human embryos.

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.

I hope my grand-kids will have natural green or purple hair. Just like in the mangas.
I hope my grand-kids will have natural green or purple hair. Just like in the mangas.
This is a wonderful advancement for prospective parents who suffer from genetic traits that give rise to birth defects. I hope the FDA soon allows these techniques to be used in the US, without increasing their price above what the market will bear.
This is a wonderful advancement for prospective parents who suffer from genetic traits that give rise to birth defects. I hope the FDA soon allows these techniques to be used in the US without increasing their price above what the market will bear.
I hope my grand-kids will have natural green or purple hair. Just like in the mangas.
This is a wonderful advancement for prospective parents who suffer from genetic traits that give rise to birth defects. I hope the FDA soon allows these techniques to be used in the US, without increasing their price above what the market will bear.