The world’s first gene detection platform made up entirely from self-assembled DNA nanostructures has been made. The other interesting aspect is to generalize the techniques used to rapidly create 100 trillion reactive and functional DNA components with easily readable results. If the attached differentiated labels could be rapidly scanned on mass then one could get a clear reading of the molecular composition of a solution. This method will allow for the barcoding of individual molecules for easy identification and analysis.
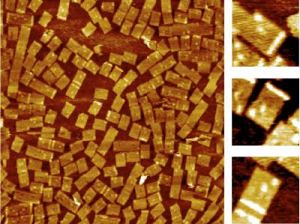

Left is an AFM image of DNA nanoarrays bound to their RNA targets at 1500nm x 1500nm scale. On the right (10 times magnification) show the barcode (white dots) that identifies the nanoarray and the RNA hybridization signal on the DNA nanoarray (white bar). Credit: Yonggang Ke
Hao Yan, Assistant Professor at Arizona State University’s Biodesign Institute, led an interdisciplinary ASU team to develop a way to use structural DNA nanotechnology to target the chemical messengers of genes, called RNA.
A recent breakthrough of making spatially addressable DNA nanoarrays came from Paul Rothemund’s work on scaffolded DNA origami, a method in which a long, single-stranded viral DNA scaffold can be folded and stapled by a large number of short synthetic “helper strands” into nanostructures that display complex patterns.
“But the potential of structural DNA nanotechnology in biological applications has been underestimated, and if we look at the process of DNA self-assembly, you will be amazed that trillions of DNA nanostructures can form simultaneously in a solution of few microliters, and very importantly, they are biocompatible and water soluble,” said Yan.
“In this work, we developed a water soluble nanoarray that can take advantage of the DNA self-assembling process and also have benefits that the macroscopic DNA microchip arrays do not have,” said Yan. “The arrays themselves are reagents, instead of solid surface chips.”
Yan refers to the self-assembled DNA nanoarrays as nucleic acid probe tiles, which look like a nanosized postage stamp. In a single step, the M13 scaffold system can churn out as many as 100 trillion of the tiles with close to 100 percent yield.
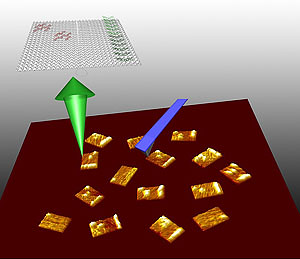
Yan’s team designed three different DNA probe tiles to detect three different RNA genes along with a bar code index to tell the tiles apart from each other. “Each probe can be distinguished by its own bar code, so we mixed them together in one solution and we used this for multiplex detection,” said Yan. The group uses a powerful instrument, atomic force microscopy (AFM), which allows the researchers to image the tiles at the single molecule level.
On the surface of each DNA probe tile is a dangling single stranded piece of DNA that can bind to the RNA target of interest. “Each probe actually contains two half probes, so when the target RNA comes in, it will hybridize to the half probes and turn the single stranded dangling probes into a stiff structure,” said Yan. “When it is stiffened, it will be sensed by the atomic force microscope cantilever, and you can see a bright line, which is a height increase. The result is a mechanical, label-free detection.”
The technology is able to detect minute quantities of RNA. “Since the DNA-RNA hybridization has such a strong affinity, in principle, a single molecule would be able to hybridize to the probe tile,” said Yan.
FURTHER READING
Hao Yan’s lab

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.

