New Scientist – It has been long known that DNA can be altered “epigenetically” – where changes occur without altering the sequence of DNA but leave chemical marks on genes that dictate how active they are by adding chemical methyl groups that silence genes, for example. Numerous environmental factors, such as stress and smoking, have been shown to influence these epigenetic marks.
Now, researchers have discovered that messenger RNA, the mirror-image copy of DNA from which all proteins are manufactured, can be methylated too.
Jaffrey’s team found that around a fifth of the RNA produced in cells from rat brains and human kidneys contained methylated versions of adenosine, one of the four building blocks of our genetic code. “It was exciting to find that 20 per cent had methyl groups, so it must be a pretty fundamental regulatory mechanism,” says Jaffrey.
Separate analyses of assorted rat tissues demonstrated that the methylated RNA was concentrated in the brain, liver and kidneys. Also, samples from rat embryos showed that concentrations rose 70-fold in the brain as it reached the final stages of growth, therefore they are likely to play a fundamental role in development.
The team also discovered that the methyl groups are stripped off the RNA by an enzyme linked with obesity. The enzyme is made by a gene called FTO, one variant of which raises the risk of obesity by 70 per cent. People with an overactive copy of the gene are most at risk, suggesting that stripping the methyl groups from RNA might somehow alter our metabolism.
The researchers found that methylated adenosine tended to cluster close to the point on the RNA strand where protein manufacture reaches completion, and on regions where other proteins bind to the strand to alter or halt production. The suggestion is that methylation may therefore dictate how much protein gets made, and when. “It’s not changing what would be made, but it might govern how much and when it’s made,” says Jaffrey. This, he says, could in turn have a big impact on a multitude of physiological processes and disease.
Technical Highlights
* m6A is a widespread RNA modification in many tissues with high levels in the brain
* MeRIP-Seq identifies m6A in 7,913 genes encoding both coding and noncoding RNAs
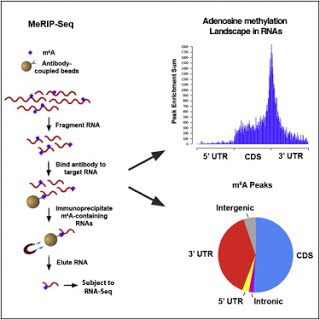
* m6A is enriched near stop codons and within 3′ UTRs in both mouse and human mRNAs
* The transcriptome-wide landscape of m6A provides important insights into m6A function
Technical Summary
Methylation of the N6 position of adenosine (m6A) is a posttranscriptional modification of RNA with poorly understood prevalence and physiological relevance. The recent discovery that FTO, an obesity risk gene, encodes an m6A demethylase implicates m6A as an important regulator of physiological processes. Here, we present a method for transcriptome-wide m6A localization, which combines m6A-specific methylated RNA immunoprecipitation with next-generation sequencing (MeRIP-Seq). We use this method to identify mRNAs of 7,676 mammalian genes that contain m6A, indicating that m6A is a common base modification of mRNA. The m6A modification exhibits tissue-specific regulation and is markedly increased throughout brain development. We find that m6A sites are enriched near stop codons and in 3′ UTRs, and we uncover an association between m6A residues and microRNA-binding sites within 3′ UTRs. These findings provide a resource for identifying transcripts that are substrates for adenosine methylation and reveal insights into the epigenetic regulation of the mammalian transcriptome.
If you liked this article, please give it a quick review on ycombinator or StumbleUpon. Thanks

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.