Chris Phoenix looks at carbohydrate strands, a new molecular building block
Cells in a multicellular organism are surrounded by a matrix of molecules called, appropriately enough, the extracellular matrix (ECM). The ECM is made up of protein and carbohydrate. It anchors the cells and provides structure to the organism. It also provides signaling for mobile cells such as immune system cells. There’s a technology being developed to build many different carbohydrate strands in an array in parallel under optical control, similar to the way an array of DNA strands can be built. The point of this – or at least, one application – is to research the way cells react with the ECM, and perhaps develop new medical sensors.
A molecular manufacturing system based on carbohydrates might start with an extension of whatever approach is being used for synthesizing the carbohydrate arrays. Then, as useful structures were built, it might be possible to mechanically protect and deprotect various sites on the carbohydrate molecules, making the chemistry simpler and more flexible
Drexler at Metamodern points out new DNA Nanotechnology
1. Ned Seeman’s lab has come the first engineered, high-quality set of 3D DNA crystals. Ned Seeman’s lab has come the first engineered, high-quality set of 3D DNA crystals This result provides a basis for organizing other components into regular 3D arrays.
We live in a macroscopic three-dimensional (3D) world, but our best description of the structure of matter is at the atomic and molecular scale. Understanding the relationship between the two scales requires a bridge from the molecular world to the macroscopic world. Connecting these two domains with atomic precision is a central goal of the natural sciences, but it requires high spatial control of the 3D structure of matter. The simplest practical route to producing precisely designed 3D macroscopic objects is to form a crystalline arrangement by self-assembly, because such a periodic array has only conceptually simple requirements: a motif that has a robust 3D structure, dominant affinity interactions between parts of the motif when it self-associates, and predictable structures for these affinity interactions. Fulfilling these three criteria to produce a 3D periodic system is not easy, but should readily be achieved with well-structured branched DNA motifs tailed by sticky ends. Complementary sticky ends associate with each other preferentially and assume the well-known B-DNA structure when they do so; the helically repeating nature of DNA facilitates the construction of a periodic array. It is essential that the directions of propagation associated with the sticky ends do not share the same plane, but extend to form a 3D arrangement of matter. Here we report the crystal structure at 4 Å resolution of a designed, self-assembled, 3D crystal based on the DNA tensegrity triangle. The data demonstrate clearly that it is possible to design and self-assemble a well-ordered macromolecular 3D crystalline lattice with precise control.

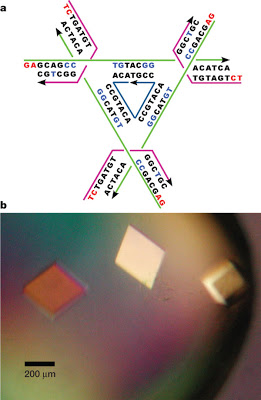
a, Schematic of the tensegrity triangle. The three unique strands are shown in magenta (strands restricted to a single junction), green (strands that extend over each edge of the tensegrity triangle) and dark blue (one unique nicked strand at the centre passing through all three junctions). Arrowheads indicate the 3′ ends of strands. Nucleotides with A-DNA-like characteristics are written in bright blue. Cohesive ends are shown in red letters. b, An optical image of crystals of the tensegrity triangle. The rhombohedral shape of the crystals and the scale are visible.
4 pages of supplemental information
2. Paul Rothemund work already blogged here. DNA Scaffolding.
3. Shih work on DNA Shapes was also already blogged here. DNA nanotech makes more shapes and tools.
Lipid and polymer-based technologies are poised to enter late-stage trials and possibly even reach the market in the next few years
Scientists trying to deliver siRNA need to engineer around several troublesome properties. RNA has a molecular weight that is 10 to 20 times that of a traditional small-molecule drug. And because the molecule is highly negatively charged, it typically can’t cross the similarly negatively charged plasma membranes to enter the cell. It’s no wonder naked strands of siRNA didn’t make it as a therapeutic approach.
Delivery systems for siRNA must overcome three major obstacles: getting the drug to its target in the body, coaxing it inside the cell, and releasing it. Even after all that is accomplished, companies then need to worry about safety, a major concern given the power of siRNA to turn off cellular processes.
Lipid- and polymer-based systems are the most established approaches for systemic delivery of RNAi. In the clinic, lipid nanoparticles (LNPs) have advanced the most. Alnylam Pharmaceuticals, widely acknowledged as the leader in the RNAi arena, has a liver cancer drug in Phase I trials that applies Tekmira Pharmaceuticals’ stable nucleic acid lipid particle technology. Alnylam is also conducting early-stage studies of other drugs that use its own LNP formulations.
One bright spot is that once researchers figure out how to get an LNP or polymer system into a particular tissue, the possibility opens for treating a host of diseases; in general, companies need only to change the siRNA payload.
So far, most of the success with both LNPs and polymer-based systems has been in delivering siRNA to the liver, which has leaky walls that enable the particles to slip in, explains Jon Wolff, vice president and head of research at Roche’s Madison, Wis., labs. Tumor vasculature and blood vessels are similarly “leaky,” and both are the subject of intensive drug development efforts, he adds.
Currently, the drug industry is focused on designing newer and better lipids that could enhance delivery to different kinds of tissues. As a result, scientists have become better at getting the particle inside the cell, but getting the siRNA out of the pathway to the destructive lysosome remains a major hurdle. “If the particle is unable to escape the endosome, it enters into a degradation pathway. Basically, it’s game over,” Akinc says.
More cutting-edge research and thinking will be needed to overcome the myriad obstacles to broader therapeutic use for siRNA. “One would like to say the path is clear,” MDRNA’s Polisky says, “but in reality this is still a large challenge.”
.

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.