Ad Support : Nano Technology Netbook Technology News Computer Software
UCLA researchers report in the April 30 edition of the journal Cell that they have imaged a virus structure at a resolution high enough to effectively “see” atoms, the first published instance of imaging biological complexes at such a resolution.
The research team, led by Hong Zhou, UCLA professor of microbiology, immunology and molecular genetics, used cryo-electron microscopy to image the structure at 3.3 angstroms. An angstrom is the smallest recognized division of a chemical element and is about the distance between the two hydrogen atoms in a water molecule.
This is the first study to determine an atomic resolution structure through Cryo-EM alone,” said Xing Zhang, a postdoctoral candidate in Zhou’s group and lead author of the Cell paper. “By proving the effectiveness of this microscopy technique, we have opened the door to a wide variety of biological studies.”
With traditional light microscopy, a magnified image of a sample is viewed through a lens. Some samples, however, are too small to diffract visible light (in the 500 to 800 nm range, or 5,000 to 8,000 angstroms) and therefore cannot be seen. To image objects at the sub-500 nm scale, scientists must turn to other tools, such as atomic force microscopes, which use an atomically thin tip to generate an image by probing a surface, in much the same way a blind person reads by touching Braille lettering.
With electron microscopy, another sub-500 nm technology, a beam of electrons is fired at a sample, passing through empty areas and bouncing off dense areas. A digital camera reads the path of the electrons passing through the sample to create a two-dimensional projection image of the sample. By repeating this process at hundreds of different angles, a computer can construct a three-dimensional image of the sample at a very high resolution.
Zhou is faculty director of the Electron Imaging Center for Nanomachines (EICN) at UCLA’s California NanoSystems Institute, which is using cryo-electron microscopy to create 3-D reconstructions of nano-machineries, nano-devices and biological nano-structures, such as viruses.
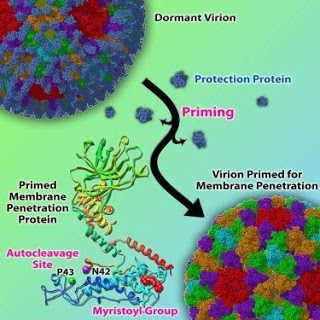
Structurally accurate 3-D reconstructions of biological complexes are possible with cryo-electron microscopy because the samples are flash frozen, which allows them to be imaged in their native environment, and the microscope operates in a vacuum, because electrons travel better in that environment. The Cell paper focused on a structural study of the aquareovirus, a non-envelope virus that causes disease in fish and shellfish, in an effort to better understand how non-envelope viruses infect host cells.
What is Cryo-EM?
Cryo-electron microscopy (also known as electron cryomicroscopy or cryoEM) is the method our lab uses to “take photographs” of viruses and other macromolecular complexes. At wikipedia there is a cryoem entry
Electron cryomicroscopy (cryo-EM or sometimes cryo-electron microscopy) is a form of electron microscopy (EM) where the sample is studied at cryogenic temperatures (generally liquid nitrogen temperatures). CryoEM is developing popularity in structural biology and protein crystallography. A version of electron cryomicroscopy is cryo-electron tomography (CET) where a 3D reconstruction of a sample is created from tilted 2D images, again at cryogenic temperatures (either liquid nitrogen or helium).
The following is an abbreviated, layman’s terms explaination of how cryoEM works.
Preparation
First, we must prepare the specimen for studying with the electron microscope. For the purposes of this example, let’s pretend we’re trying to image a virus. We grow it to a concentrated number (or high titer), isolate it, and purify it. Then, when we have a good sample, we place a drop containing thousands of virions onto a thin film which is then quickly frozen to the temperature of liquid nitrogen in order to protect and preserve the specimen during observation.
Imaging
When the sample is ready, we can start shooting electrons at it. Cryo-EM uses a very low dose of electrons (about 1-10 electrons per square angstrom) so that the biological sample is not damaged during the study. The electrons pass through empty areas and are bounced or refracted from dense areas. Please note that although the lenses in the image to the left look an awful lot like a magnifying glass, electron microscopes actually use magnetic coils to “magnify” and focus the electrons.
Final Results
When imaging is complete, we end up a two-dimensional image similar to the one you see in the diagram. To render these flat images into a three-dimensional model, we must use computerized 3-D data merging.
Further information including videos at this link
If you liked this article, please give it a quick review on Reddit, or StumbleUpon. Thanks
Supporting Advertising
Business Success
How to Make Money
Executive Jobs
Paid Surveys
Thank You

Brian Wang is a Futurist Thought Leader and a popular Science blogger with 1 million readers per month. His blog Nextbigfuture.com is ranked #1 Science News Blog. It covers many disruptive technology and trends including Space, Robotics, Artificial Intelligence, Medicine, Anti-aging Biotechnology, and Nanotechnology.
Known for identifying cutting edge technologies, he is currently a Co-Founder of a startup and fundraiser for high potential early-stage companies. He is the Head of Research for Allocations for deep technology investments and an Angel Investor at Space Angels.
A frequent speaker at corporations, he has been a TEDx speaker, a Singularity University speaker and guest at numerous interviews for radio and podcasts. He is open to public speaking and advising engagements.